
Action and Regulation of Bromodomain-Containing Proteins
People working on this project:
Brian Smith, Mike Olp, Chris Goetz, Brad Julga
Recent estimates indicate that the human genome only contains ~19,000 protein encoding genes, or slightly less than the nematode worm genome. Protein post-translational modification is largely responsible for the complexity arising from the small number of genes that humans possess. The Smith lab focuses on the post-translational modification of critical charged and nucleophilic residues by lysine acylation and cysteine S-nitrosation.
Lysine acylation
Post-translational modification of histones, transcription factors, and other nuclear proteins forms the basis of epigenetics and regulates transcription through unknown mechanisms. The combinatorial effect of these post-translational modifications underlies the “histone language” that is interpreted by three broad protein classes: “writers”, “readers”, and “erasers”. Lysine residues are particularly abundant targets of epigenetic regulation and are modified by an astounding array of modifications including methylation, ubiquitinylation, sumoylation, ADP-ribosylation, oxidation, and acylation. One focus of my laboratory will be on the mechanism and regulation of the bromodomain family of proteins that read sites of epigenetic lysine acylation.
Bromodomain reading of the acyl-lysine histone language
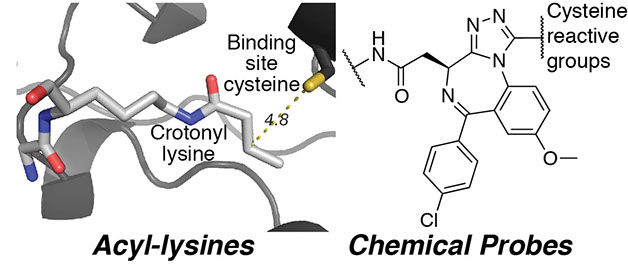
Bromodomain inhibitors are being developed to treat leukemia, lymphoma, and inflammation. Although bromodomains were originally described as readers of acetyl-lysine, many other histone acyl-lysine modifications were recently discovered. The distinct functions of these unique acyl-lysines are unknown; these functions will be elucidated in this project. The lab also aims to develop chemical probes allowing discovery of bromodomain drug targets and to profile bromodomain inhibitors.

© 2017 Brian Smith Lab
Department of Biochemistry
Medical College of Wisconsin
8701 Watertown Plank Rd,
Milwaukee, WI, 53226 | (414) 955-5659
